Same as gene_specificity_scores.long_, but filtered to significantly specific genes (see methods). Peak_specificity_scores_sigsites.long_format.rds Same as peak_specificity_scores.long_, but filtered to significantly specific sites (see methods). site column replaced with gene_short_name.

Same as peak_specificity_scores.long_, but gene-level specificity calculated from activity scores.
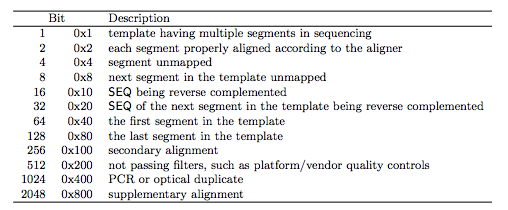

The distribution family used by monocle (binomialff in this case)īeta derived from the model returned by monocle. Status of successful test completion reported by monocle (OK for all in this case) Note that files contain one entry per combination of cluster, subset_cluster, and peak/gene_short_name. The following columns are provided in each DA test file. Metadata for peaks-intersected TSS pairs in TSV format Provided matrices would need to be subsetted to match this set of cells if using this metadata. These very low frequency labels are often not cell types expected their respective tissues and could be due to slight imperfections in clustering, for example. Same as cell_metadata.txt, but removes cells in each tissue belonging to a cell_label that accounts for less than 0.5% of the cells in that tissue. id: combined ID for major + iterative cluster assignment.subset_tsne2: t-SNE2 coordinate in iterative t-SNE.subset_tsne1: t-SNE1 coordinate in iterative t-SNE.
tsne_2: t-SNE2 coordinate in initial t-SNE.tsne_1: t-SNE1 coordinate in initial t-SNE.subset_cluster: cluster assignment in iterative t-SNE space.cluster: cluster assignment in initial t-SNE.tissue_replicate: same as tissue, but each replicate has a unique id.tissue: the tissue that this cell originated from.cell: cell barcode (combined and corrected).Gene names (common names columns) of binarized gene activity score matrix.īinarized gene activity score matrix in RDS format. Quantitative gene activity score matrix in RDS format.īinarized gene activity score matrix in matrix market format.Ĭell IDs (columns) of binarized gene activity score matrix. Gene names (common names columns) of quantitative gene activity score matrix. Quantitative gene activity score matrix in matrix market format.Ĭell IDs (columns) of quantitative gene activity score matrix. Quantitative scores provided here are not normalized by size factors, so you may apply size factor normalization to these values if needed. TFIDF normalized peak by cell matrix in RDS format. TFIDF normalized peak by cell matrix in matrix market format.
RDS format that can be read into R directly with readRDS function.īinarized peak by cell matrix in matrix market format.īinarized peak by cell matrix in RDS format. txt files containing peak and cell IDs that correspond to the rows and columns of the matrix, respectively. Same as format generated by 10X Genomics cellranger pipeline (matrix market format).mtx files are provided with. In general, we provide two formats for all matrices: File Type
